Many drug and antibody discovery pathways focus on intricately folded cell membrane proteins. When molecules of a drug candidate bind to these proteins, like a key going into a lock, they trigger chemical cascades that alter cellular behavior. Understanding how proteins fold and move is therefore essential for developing drugs that interact well with their targets.
Artificial intelligence (AI) is a very useful tool to generate novel protein structures, but most systems—including Google DeepMind’s AlphaFold—focus on producing static “snapshots” of proteins. Subtle rearrangements of atoms in structures called side chains, which influence a protein’s interactions with other molecules, are not captured.
Now, scientists in EPFL’s School of Life Sciences have teamed up with data processing experts in the School of Engineering to solve this problem.
Researchers led by Patrick Barth of the Laboratory of Protein and Cell Engineering (LPCE) and Pierre Vandergheynst of the Signal Processing Laboratory (LTS2) have developed an AI-based generative framework called Latent Diffusion for Full Protein Generation (LD-FPG), which produces complete, all-atom structural ensembles of proteins and their movements.
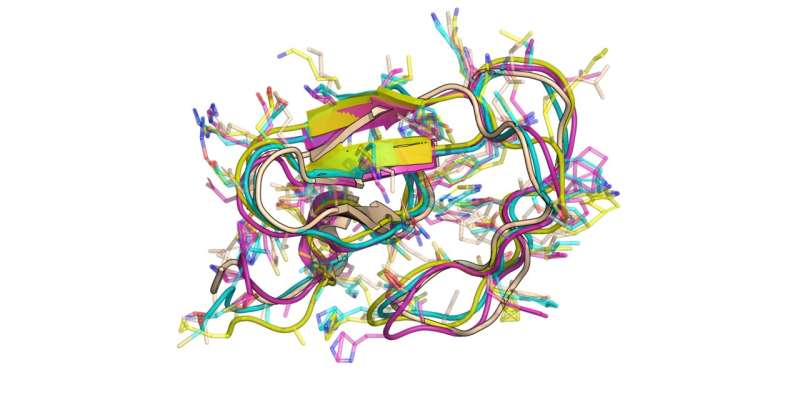
“Proteins are like tiny machines that dance and switch on and off to work, but generating this ‘movie’ in full detail has been an unsolved challenge,” says LPCE researcher Aditya Sengar.
“Our LD-FPG framework is the first to do this. Instead of trying to predict the exact coordinates of atoms in space, our model learns a low-dimensional map of the protein’s shape changes. This conceptual shift is what makes generating all-atom dynamics possible.”
The new framework can notably generate the full range of motion for complex drug targets like G-protein coupled receptors (GPCRs), a focus of the global drug development industry.
“LD-FPG opens the door to designing new medicines that target a protein’s dynamic behavior, not just its shape. Our work represents a new paradigm for computational biology, and a meaningful step forward at the interface of AI and structural biology,” says Barth.
The work has been published in the Proceedings of NeurIPS 2025.
Capturing a protein’s dance
Because systems like AlphaFold use AI to predict the spatial position of every atom in a protein, they require vast amounts of computing power and biology and computer science expertise.
LD-FPG simplifies this problem using something called a graph neural network (GNN). The GNN treats each protein like a mathematical graph, where atoms represent “nodes” and the bonds between them represent “edges.” Using this low-level representation, it essentially compresses protein structure data into a simplified, or latent, map.
Next, an AI model studies this map and “learns” the representations of the protein’s structure and movements. Once trained, the model generates latent data for entirely new structures. Finally, these simplified data are converted back into high-resolution proteins—complete with side chains and dynamic movements.
In one experiment, the team generated high-fidelity, dynamic representations of the dopamine D2 receptor in both its active and inactive states. This protein detects the neurotransmitter dopamine and controls key cellular responses, making it one of the most-studied GPCRs. The researchers have published this dataset with open access to facilitate further research.
“In addition to enhancing biological understanding, we believe our work will help improve virtual screening processes for proteins, which currently involve a lot of trial and error, thereby accelerating drug discovery,” Sengar says.
Going forward, the team aims to streamline the AI framework for even greater accuracy and realism, and to enable it to model larger proteins. But Vandergheynst emphasizes that high-quality data will remain the bedrock of success. “Many assume that feeding massive datasets to AI models will automatically solve scientific problems or replace researchers.
“However, much of that data is noisy or poorly evaluated. We need human scientists to produce the clean data and rigorous benchmarks AI requires, much like we need journalists to safeguard against disinformation.”