Titanium particles may explain why antibiotics fail against dental implant infections
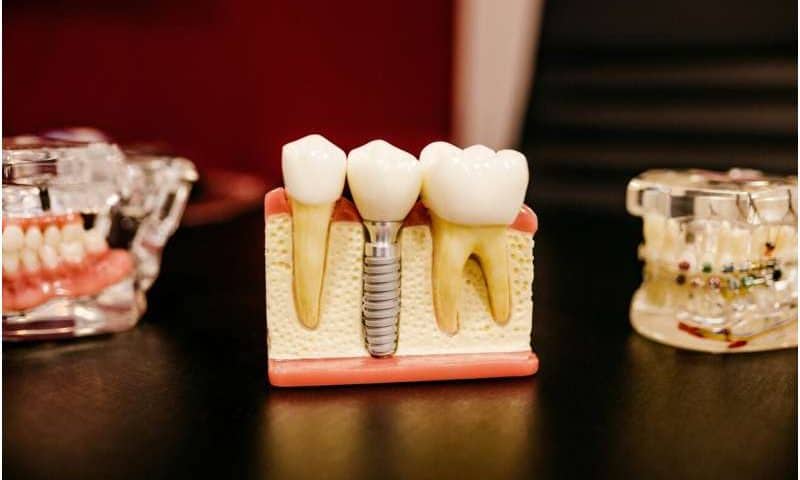
Dental implants have given tens of millions of people something dentures never could: a full set of fixed and fully functioning teeth. Unfortunately, 10% to 20% of implant patients eventually experience an aggressive jawbone infection called peri-implantitis. Antibiotics usually fail to stop the infection for reasons that researchers have not understood until now.
Study uncovers a surprising culprit
A recent study in PNAS Nexus by researchers with the Rutgers School of Dental Medicine has found that bacteria corrode implants, causing them to shed microscopic titanium particles into the surrounding tissue. Those particles hijack the immune cells sent to clear the infection and lock them into a state of inflammation that destroys the jawbone they are supposed to protect.
Working with human tissue samples, cultured human immune cells and a genetically engineered mouse model, the team pinpointed a specific calcium channel in the body’s bacteria-eating macrophages that the titanium particles activate. Switching that channel off in mice prevented the disease. The result is the first credible drug target for a condition that affects up to one in five implant recipients and costs the global health system more than a billion dollars a year.
“For the first time, we show why all the antibiotic treatments that work around teeth do not work around implants,” said Georgios Kotsakis, the study’s senior author and the assistant dean for clinical research at the dental school. “Now that we know the cause, we can start developing therapeutics.”
Why implants behave differently than teeth
Peri-implantitis has long been a puzzle because it initially looks like its counterpart in natural teeth, which is called periodontitis and begins with the same oral bacteria. In patients with natural teeth, antibiotics and routine cleaning resolve the infection. In patients with implants, the same drugs against the same bacteria succeed less than half the time, while the bone underneath continues to disappear.
Most research over the past 20 years has focused on the bacteria. Members of Kotsakis’s lab took a different approach and began looking at the implants. Bacteria living on the implant surface produce acidic biofilms that slowly corrode the titanium, releasing billions of particles smaller than a red blood cell. The same shedding can occur during routine cleaning, especially with instruments that dentists typically use on natural teeth.
Inside the gum, those particles get coated with a bacterial toxin called lipopolysaccharide. To the immune system, they suddenly look like enormous, indigestible bacteria. Macrophages, a type of white blood cell that surrounds and kills microorganisms, engulf them but cannot digest metal. The cells become trapped in a hyperinflammatory state, pumping out signaling molecules including interleukin-1 beta, an inflammatory protein also implicated in rheumatoid arthritis and Alzheimer’s disease.
How titanium particles hijack immunity
That inflammation eats away at bone. Worse, the immune cells lose their ability to deal with the original infection. In the lab, macrophages exposed to titanium particles took up less than half as many bacteria as unexposed cells.
“These particles are little magnets that attract the bacterial toxin, and they hijack the immune system, preventing it from clearing bacteria,” said Kotsakis. “You have a perfect storm that defies antibiotics.”
Team members traced the cascade to a calcium channel (a specialized, pore-forming protein structure within cell membranes) called TRPC1. In mice engineered without it, the immune cells handled the same titanium-plus-bacteria challenge normally: Abscesses were dramatically smaller, inflammatory cytokines dropped, and bacterial clearance was restored.
New treatment avenues and safer cleanings
Members of Kotsakis’s group are now testing drug candidates that target the same pathway in human cells.
For people who already have implants, the most useful finding may be a quieter one. The strongest known protective factor is regular professional cleaning, but the kind of cleaning matters. Until roughly a decade ago, many dentists scraped implants with the metal scalers used on teeth, a method the Rutgers lab and others have shown can itself corrode the implant and accelerate the disease. Nonabrasive techniques are now standard.