The hippocampus is an important brain region known to support various cognitive (i.e., mental) processes, including the encoding and retrieval of memories, learning, decision-making and the regulation of emotional states. While extensive research has tried to delineate the structure, functions and organization of the hippocampus, the cell types contained within it and their connections with other neurons have not yet been fully mapped out.
Over the past decades, available methods for studying cell subpopulations, the expressions of genes within them and their connectivity have become increasingly advanced. One of these methods, known as spatially resolved transcriptomics, works by measuring the expression of genes in cells while preserving their arrangement in space. Another called single-nucleus RNA-sequencing (snRNA-seq), allows scientists to examine RNA molecules inside individual cell nuclei to detect differences between them and categorize cells into different subtypes.
Researchers at Johns Hopkins Bloomberg School of Public Health, the Lieber Institute for Brain Development and Johns Hopkins School of Medicine recently used a combination of these two experimental techniques to examine cells in tissue extracted from the hippocampus. Their paper, published in Nature Neuroscience, introduces a comprehensive molecular atlas of the hippocampus that maps different cell subtypes and their organization.
“Cell types in the hippocampus with unique morphology, physiology and connectivity serve specialized functions associated with cognition and mood,” wrote Jaqueline R. Thompson, Erik D. Nelson and their colleagues in their paper. “These cell types are spatially organized, necessitating molecular profiling strategies that retain cytoarchitectural organization. We generated SRT and snRNA-seq data from anterior human hippocampus in ten adult neurotypical donors.”
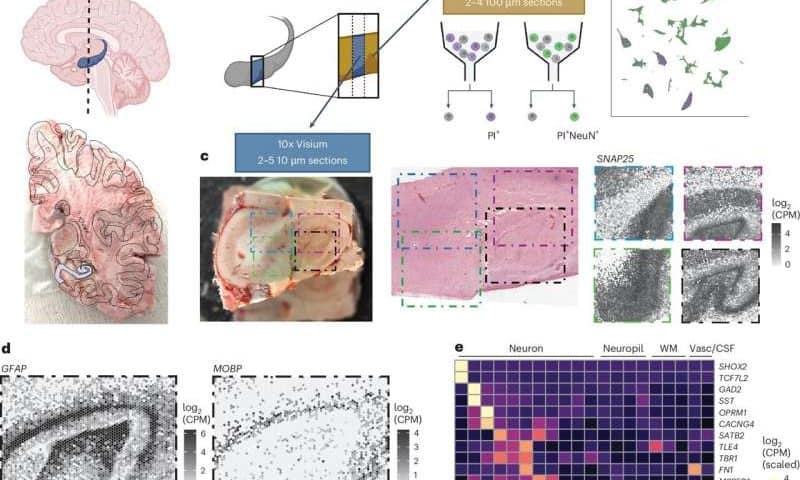
As part of their study, Thompson, Nelson and their colleagues analyzed hippocampal tissue that was extracted from the brains of 10 humans with no reported brain anomalies. They then derived SRT and sRNA-seq data for cells in the tissues they examined and used computational tools to combine the two types of data.
“Using non-negative matrix factorization (NMF) and label transfer, we integrated this data by defining gene expression patterns within the snRNA-seq data and then inferring expression in the SRT data,” wrote the authors. “These patterns captured transcriptional variation across neuronal cell types and indicated spatial organization of excitatory and inhibitory postsynaptic specializations.”
Through their analysis of hippocampal tissue, the researchers were able to map the locations of different cell types in the hippocampus. They then compared their results with those attained when examining the mouse hippocampus.
“Leveraging the NMF and label transfer approach with rodent datasets, we identified putative patterns of activity-dependent transcription and circuit connectivity in the human SRT dataset,” wrote Thompson, Nelson and their colleagues. “Finally, we characterized the spatial organization of NMF patterns corresponding to pyramidal neurons and identified regionally specific snRNA-seq clusters of the retrohippocampus, subiculum and presubiculum.”
Notably, the molecular atlas created by Thompson, Nelson and their colleagues, along with the raw data it is based on, can be accessed online via an interactive web application. In the future, it could serve as a reference for other teams conducting medicine and neuroscience research focusing on the hippocampus.