Since addressing the COVID-19 pandemic, mRNA vaccines have risen as a promising tool for treating a broad range of diseases, such as cancer and genetic disorders. To be effective, foreign mRNA in these vaccines must evade the body’s natural defenses to enter cells and synthesize proteins. However, mechanisms governing mRNA cellular regulation remain poorly understood.
In a new study published in Science titled, “Exogenous RNA surveillance by proton-sensing TRIM25,” researchers from the Institute for Basic Science (IBS) in South Korea, have identified cellular factors that regulate the delivery and stability of mRNA via a pH-dependent surveillance system to expand the potential of mRNA therapeutics.
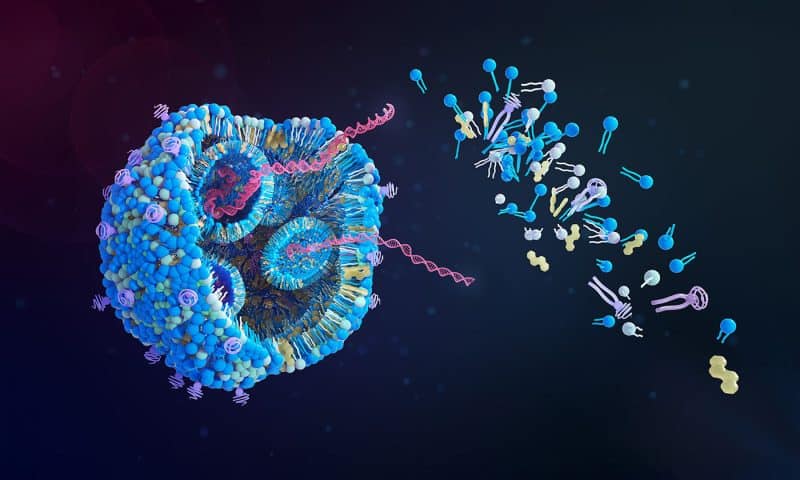
The study, led by Narry Kim, PhD, professor at Seoul National University and director of the Center for RNA Research at the IBS, used genome-wide CRISPR screens targeting 19,114 genes to pinpoint key factors that facilitate the cellular uptake and stability of exogenous mRNAs. mRNAs encoding Green Fluorescent Protein (GFP) were delivered to knockout cells using lipid nanoparticles (LNPs). Cells with high or low GFP fluorescence were collected and sequenced to uncover cellular regulators of LNP-encapsulated mRNAs.
To elucidate mRNA stability, the study identified that the binding of TRIM25, a cellular defense protein, to exogenous mRNAs induced rapid degradation to prevent function. mRNA containing the modification, N1-methylpseudouridine (m1Ψ), evaded TRIM25 binding to enhance mRNA stability.
Notably, the 2023 Nobel Prize in Physiology or Medicine was awarded to Drew Weissman and Katalin Karikó for the discovery of nucleoside base modifications that enabled the development of effective mRNA vaccines against COVID-19. The results from this study provide a molecular mechanism by which clinically relevant m1Ψ enhances the therapeutic potential of mRNA-based treatments.
The researchers also demonstrated that protons played a key role as immune signaling molecules. TRIM25’s RNA binding affinity increased even at mildly acidic pH, which indicated activation by protons released from ruptured endosomes. The authors noted that TRIM25 functions across multiple human and mouse cell types, suggesting a widespread and conserved surveillance mechanism.
Additional mRNA regulatory factors identified by the study included heparan sulfate (HSPG), a glycoprotein on the cell surface, and vascular ATPase, which mediate mRNA uptake and endosomal escape, respectively. Interfering with HSPGs reduced mRNA internalization while inhibition of vascular ATPase blocked endosomal escape of LNP-mRNA.
Looking ahead, the authors stated that this comprehensive map of cellular pathways regulating LNP-mRNAs offers novel insights into RNA immunity to develop more effective therapeutics.
“Understanding how cells respond to mRNA vaccines is key to improving mRNA therapeutics. To develop effective RNA treatments, we need to find ways to bypass the cellular defense mechanisms and harness the endosomal system effectively,” said Kim.
Other mRNA therapeutics projects explored by the Kim lab include identifying viral RNAs with potent regulatory activities through high-throughput sequencing-based screens, the effect of mRNA tail modifications on stability, and the role of microRNA in regulating cell function, including differentiation, proliferation, cell death, and energy metabolism.